Radial & Angular Distribution Function
Radial and angular distribution function (RDF & ADF) generators have been
implemented in the FOX.MultiMolecule class.
The radial distribution function, or pair correlation function, describes how
the particale density in a system varies as a function of distance from a
reference particle. The herein implemented function is designed for
constructing RDFs between all possible (user-defined) atom-pairs.
Given a trajectory, mol, stored as a FOX.MultiMolecule instance, the RDF
can be calculated with the following
command: rdf = mol.init_rdf(atom_subset=None, low_mem=False).
The resulting rdf is a Pandas dataframe, an object which is effectively a
hybrid between a dictionary and a NumPy array.
A slower, but more memory efficient, method of RDF construction can be enabled
with low_mem=True, causing the script to only store the distance matrix
of a single molecule in memory at once. If low_mem=False, all distance
matrices are stored in memory simultaneously, speeding up the calculation
but also introducing an additional linear scaling of memory with respect to
the number of molecules.
Note: Due to larger size of angle matrices it is recommended to use
low_mem=False when generating ADFs.
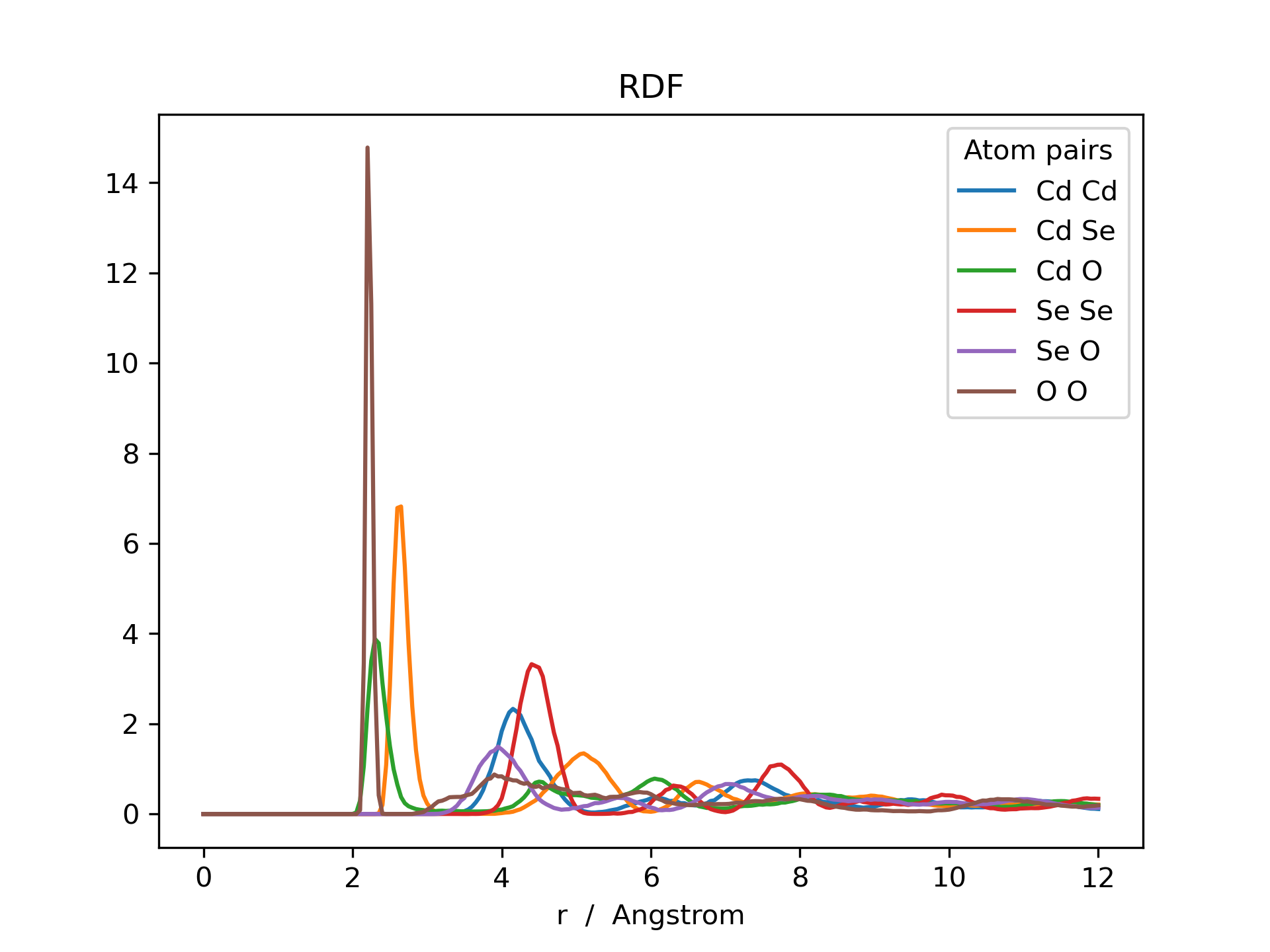
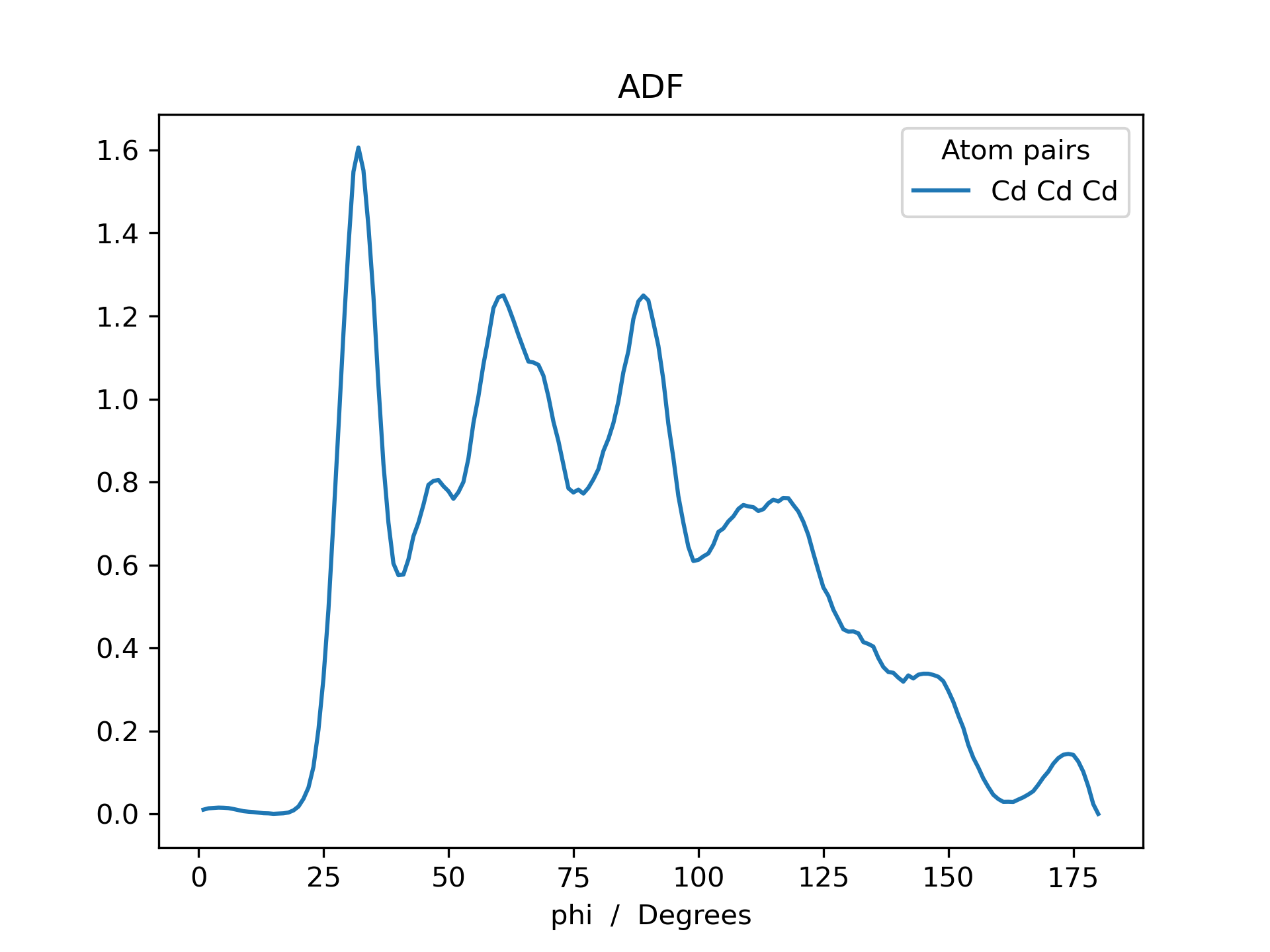
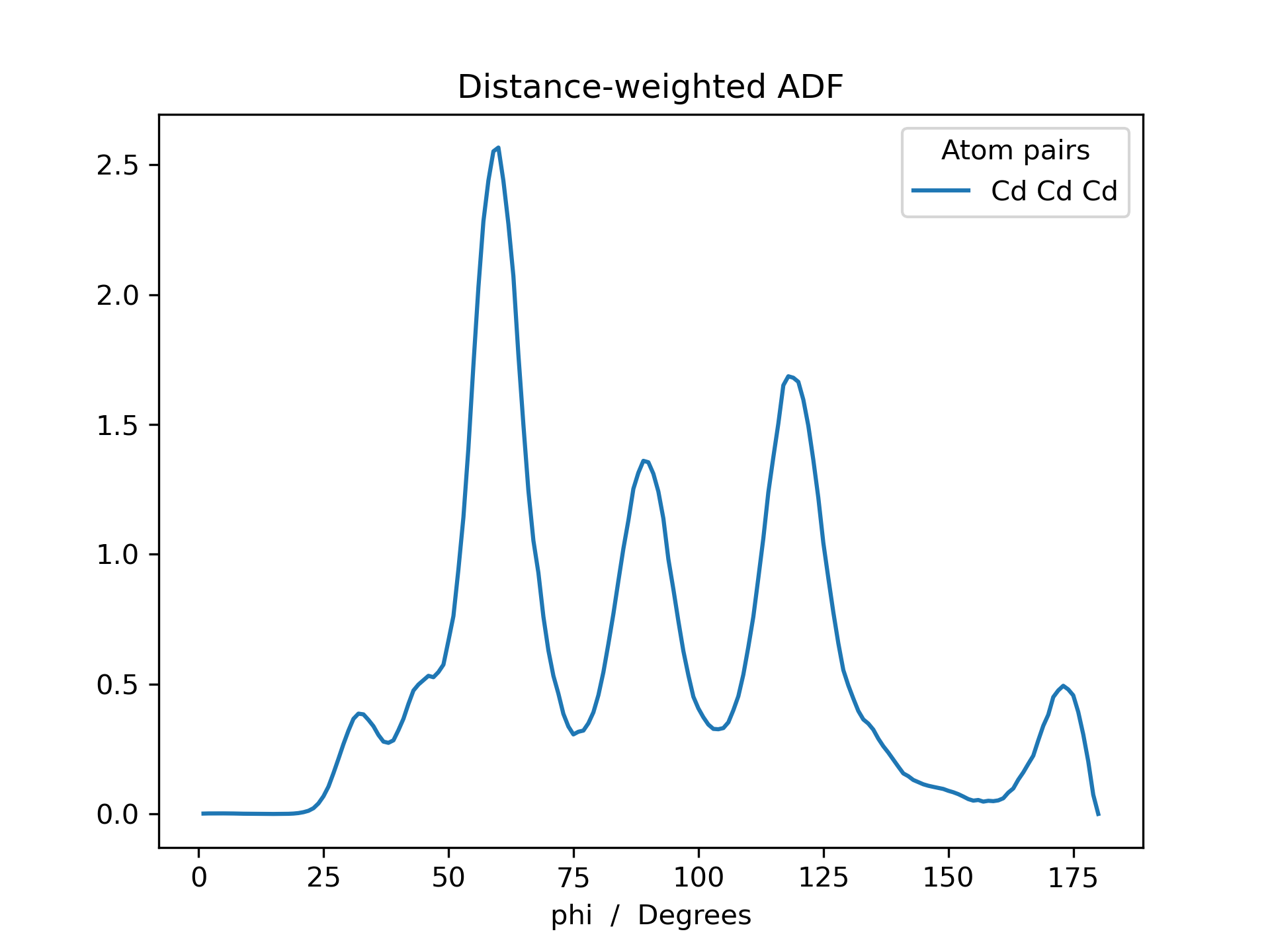
Below is an example RDF and ADF of a CdSe quantum dot pacified with formate ligands. The RDF is printed for all possible combinations of cadmium, selenium and oxygen (Cd_Cd, Cd_Se, Cd_O, Se_Se, Se_O and O_O).
>>> from FOX import MultiMolecule, example_xyz
>>> mol = MultiMolecule.from_xyz(example_xyz)
# Default weight: np.exp(-r)
>>> rdf = mol.init_rdf(atom_subset=('Cd', 'Se', 'O'))
>>> adf = mol.init_adf(r_max=8, weight=None, atom_subset=('Cd',))
>>> adf_weighted = mol.init_adf(r_max=8, atom_subset=('Cd',))
>>> rdf.plot(title='RDF')
>>> adf.plot(title='ADF')
>>> adf_weighted.plot(title='Distance-weighted ADF')
One can take into account a systems periodicity by settings the molecules’
lattice vectors and specifying the axes along which
the system is periodic.
The lattice vectors can be provided in one of two formats:
A \((3, 3)\) matrix.
A \((N_{mol}, 3, 3)\)-shaped tensor if they vary across the trajectory.
>>> from FOX import MultiMolecule
>>> import numpy as np
>>> lattice = np.array(...)
>>> mol = MultiMolecule.from_xyz(...)
>>> mol.lattice = lattice
# Periodic along the x, y and/or z axes
>>> rdf = mol.init_rdf(atom_subset=('Cd', 'Se', 'O'), periodic="xy")
>>> adf = mol.init_adf(r_max=8, atom_subset=('Cd',), periodic="xyz")
API
- MultiMolecule.init_rdf(mol_subset=None, atom_subset=None, *, dr=0.05, r_max=12.0, periodic=None, atom_pairs=None)[source]
Initialize the calculation of radial distribution functions (RDFs).
RDFs are calculated for all possible atom-pairs in atom_subset and returned as a dataframe.
- Parameters
mol_subset (
slice, optional) – Perform the calculation on a subset of molecules in this instance, as determined by their moleculair index. Include all \(m\) molecules in this instance ifNone.atom_subset (
Sequence[str], optional) – Perform the calculation on a subset of atoms in this instance, as determined by their atomic index or atomic symbol. Include all \(n\) atoms per molecule in this instance ifNone.dr (
float) – The integration step-size in Ångström, i.e. the distance between concentric spheres.r_max (
float) – The maximum to be evaluated interatomic distance in Ångström.periodic (
str, optional) – If specified, correct for the systems periodicity ifself.lattice is not None. Accepts"x","y"and/or"z".atom_pairs (
Iterable[tuple[str, str]]) – An explicit list of atom-pairs for the to-be calculated distances. Note that atom_pairs and atom_subset are mutually exclusive.
- Returns
A dataframe of radial distribution functions, averaged over all conformations in xyz_array. Keys are of the form: at_symbol1 + ‘ ‘ + at_symbol2 (e.g.
"Cd Cd"). Radii are used as index.- Return type
- MultiMolecule.init_adf(mol_subset=None, atom_subset=None, *, r_max=8.0, weight=<function neg_exp>, periodic=None, atom_pairs=None)[source]
Initialize the calculation of distance-weighted angular distribution functions (ADFs).
ADFs are calculated for all possible atom-pairs in atom_subset and returned as a dataframe.
- Parameters
mol_subset (
slice, optional) – Perform the calculation on a subset of molecules in this instance, as determined by their moleculair index. Include all \(m\) molecules in this instance ifNone.atom_subset (
Sequence[str], optional) – Perform the calculation on a subset of atoms in this instance, as determined by their atomic index or atomic symbol. Include all \(n\) atoms per molecule in this instance ifNone.r_max (
float) – The maximum inter-atomic distance (in Angstrom) for which angles are constructed. The distance cuttoff can be disabled by settings this value tonp.inf,"np.inf"or"inf".weight (
Callable[[np.ndarray], np.ndarray], optional) – A callable for creating a weighting factor from inter-atomic distances. The callable should take an array as input and return an array. Given an angle \(\phi_{ijk}\), to the distance \(r_{ijk}\) is defined as \(max[r_{ij}, r_{jk}]\). Set toNoneto disable distance weighting.periodic (
str, optional) – If specified, correct for the systems periodicity ifself.lattice is not None. Accepts"x","y"and/or"z".atom_pairs (
Iterable[tuple[str, str, str]]) – An explicit list of atom-triples for the to-be calculated angles. Note that atom_pairs and atom_subset are mutually exclusive.
- Returns
A dataframe of angular distribution functions, averaged over all conformations in this instance.
- Return type
Note
Disabling the distance cuttoff is strongly recommended (i.e. it is faster) for large values of r_max. As a rough guideline,
r_max="inf"is roughly as fast asr_max=15.0(though this is, of course, system dependant).Note
The ADF construction will be conducted in parralel if the DASK package is installed. DASK can be installed, via anaconda, with the following command:
conda install -n FOX -y -c conda-forge dask.
- FOX.recipes.time_resolved_rdf(mol, start=0, stop=None, step=500, **kwargs)[source]
Calculate the time-resolved radial distribution function (RDF).
Examples
>>> from FOX import MultiMolecule, example_xyz >>> from FOX.recipes import time_resolved_rdf # Calculate each RDF over the course of 500 frames >>> time_step = 500 >>> mol = MultiMolecule.from_xyz(example_xyz) >>> rdf_list = time_resolved_rdf( ... mol, step=time_step, atom_subset=['Cd', 'Se'] ... )
- Parameters
mol (
MultiMolecule) – The trajectory in question.start (
int) – The initial frame.stop (
int, optional) – The final frame. Set toNoneto iterate over all frames.step (
int) – The number of frames per individual RDF. Note that lower step values will result in increased numerical noise.**kwargs (
Any) – Further keyword arguments forinit_rdf().
- Returns
A list of dataframes, each containing an RDF calculated over the course of step frames.
- Return type
See also
init_rdf()Calculate the radial distribution function.
- FOX.recipes.time_resolved_rdf(mol, start=0, stop=None, step=500, **kwargs)[source]
Calculate the time-resolved radial distribution function (RDF).
Examples
>>> from FOX import MultiMolecule, example_xyz >>> from FOX.recipes import time_resolved_rdf # Calculate each RDF over the course of 500 frames >>> time_step = 500 >>> mol = MultiMolecule.from_xyz(example_xyz) >>> rdf_list = time_resolved_rdf( ... mol, step=time_step, atom_subset=['Cd', 'Se'] ... )
- Parameters
mol (
MultiMolecule) – The trajectory in question.start (
int) – The initial frame.stop (
int, optional) – The final frame. Set toNoneto iterate over all frames.step (
int) – The number of frames per individual RDF. Note that lower step values will result in increased numerical noise.**kwargs (
Any) – Further keyword arguments forinit_rdf().
- Returns
A list of dataframes, each containing an RDF calculated over the course of step frames.
- Return type
See also
init_rdf()Calculate the radial distribution function.